Brian Yu, Harvard University, brian@cs.harvard.edu
David J. Malan, Harvard University, malan@harvard.edu
| Summary | Students write a Python program that accepts (1) a CSV file representing a DNA database and (2) a text file representing a DNA sequence. Using a combination of loops and string manipulation, and file I/O, students identify to whom the DNA sequence belongs. |
| Topics | Algorithms. Computational Biology. File I/O. Loops. Python. String Manipulation. |
| Audience | Appropriate for CS1 or higher. |
| Difficulty | This is an intermediate assignment, taking about 1 week for a CS1 or CS2 student. |
| Strengths | The assignment shows a connection between computation and biology, which works well for students from the natural sciences interested in computer science; it also demonstrates a very real-world application of algorithmic thinking and string manipulation. The assignment also offers a nice balance between exploring features of the Python programming language and also thinking algorithmically about how to compute the longest run of a particular substring. |
| Weaknesses | The computation of the longest run can be a bit tricky. Several of the easier solutions are not necessarily the most efficient solutions. |
| Dependencies | Familiarity with strings, lists, CSV files. Exposure to (but not necessarily experience with) Python. |
| Variants | More comfortable students can find more efficient algorithms for computing the longest run of a substring. |
DNA, the carrier of genetic information in living things, has been used in criminal justice for decades. But how, exactly, does DNA profiling work? Given a sequence of DNA, how can forensic investigators identify to whom it belongs?
Well, DNA is really just a sequence of molecules called nucleotides, arranged into a particular shape (a double helix). Each nucleotide of DNA contains one of four different bases: adenine (A), cytosine (C), guanine (G), or thymine (T). Every human cell has billions of these nucleotides arranged in sequence. Some portions of this sequence (i.e. genome) are the same, or at least very similar, across almost all humans, but other portions of the sequence have a higher genetic diversity and thus vary more across the population.
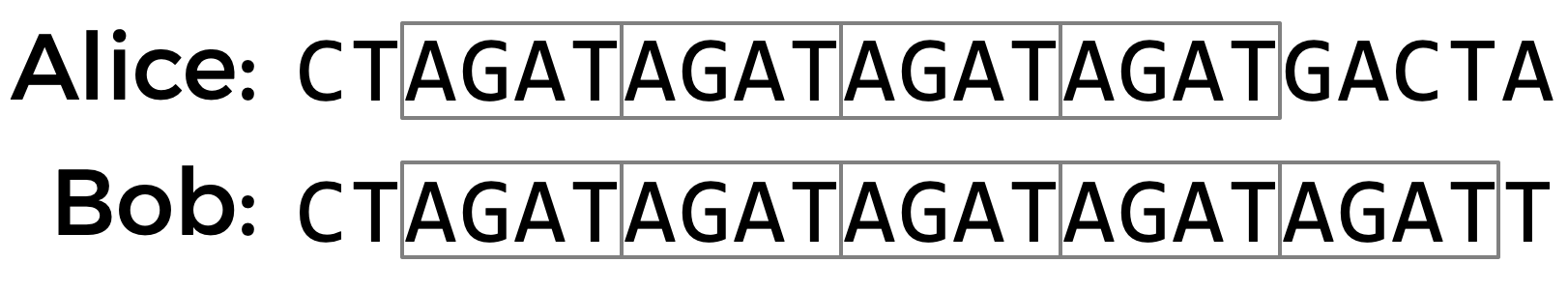
One place where DNA tends to have high genetic diversity is in Short Tandem Repeats (STRs). An STR is a short sequence of DNA bases that tends to be repeated back-to-back numerous times at specific locations in DNA. The number of times any particular STR repeats varies a lot among different people. In the DNA samples below, for example, Alice has the STR AGAT repeated four times in her DNA, while Bob has the same STR repeated five times.
Using multiple STRs, rather than just one, can improve the accuracy of DNA profiling. If the probability that two people have the same number of repeats for a single STR is 5%, and the analyst looks at 10 different STRs, then the probability that two DNA samples match purely by chance is about 1 in 1 quadrillion (assuming all STRs are independent of each other). So if two DNA samples match in the number of repeats for each of the STRs, the analyst can be pretty confident they came from the same person. CODIS, The FBI's DNA database, uses 20 different STRs as part of its DNA profiling process.
What might such a DNA database look like? Well, in its simplest form, you could imagine formatting a DNA database as a CSV file, wherein each row corresponds to an individual, and each column corresponds to a particular STR.
name,AGAT,AATG,TATC
Alice,28,42,14
Bob,17,22,19
Charlie,36,18,25
The data in the above file would suggest that Alice has the sequence AGAT repeated 28 times consecutively somewhere in her DNA, the sequence AATG repeated 42 times, and TATC repeated 14 times. Bob, meanwhile, has those same three STRs repeated 17 times, 22 times, and 19 times, respectively. And Charlie has those same three STRs repeated 36, 18, and 25 times, respectively.
So given a sequence of DNA, how might you identify to whom it belongs? Well, imagine that you looked through the DNA sequence for the longest consecutive sequence of repeated AGATs and found that the longest sequence was 17 repeats long. If you then found that the longest sequence of AATGs is 22 repeats long, and the longest sequence of TATC is 19 repeats long, that would provide pretty good evidence that the DNA was Bob's. Of course, it's also possible that once you take the counts for each of the STRs, it doesn't match anyone in your DNA database, in which case you have no match.
In practice, since analysts know on which chromosome and at which location in the DNA an STR will be found, they can localize their search to just a narrow section of DNA. But we'll ignore that detail for this problem.
Your task is to write a program that will take a sequence of DNA and a CSV file containing STR counts for a list of individuals and then output to whom the DNA (most likely) belongs.
$ python profile.py data.csv sequence.txt Alice
name,AGAT,AATG,TATC Alice,5,2,8 Bob,3,7,4 Charlie,6,1,5The following sequence should match with Alice:
AGACGGGTTACCATGACTATCTATCTATCTATCTATCTATCTATCTATCACGTACGTACGTATCGAGATAGATAGATAGATAGATCCTCGACTTCGATCGCAATGAATGCCAATAGACAAAAThe following sequence should match with Bob:
AACCCTGCGCGCGCGCGATCTATCTATCTATCTATCCAGCATTAGCTAGCATCAAGATAGATAGATGAATTTCGAAATGAATGAATGAATGAATGAATGAATGThe following sequence should match with Charlie:
CCAGATAGATAGATAGATAGATAGATGTCACAGGGATGCTGAGGGCTGCTTCGTACGTACTCCTGATTTCGGGGATCGCTGACACTAATGCGTGCGAGCGGATCGATCTCTATCTATCTATCTATCTATCCTATAGCATAGACATCCAGATAGATAGATCAnd the following sequence should have No match:
GGTACAGATGCAAAGATAGATAGATGTCGTCGAGCAATCGTTTCGATAATGAATGAATGAATGAATGAATGAATGACACACGTCGATGCTAGCGGCGGATCGTATATCTATCTATCTATCTATCAACCCCTAG